Argonne National Lab adds ‘AI supercomputer,’ boosting work of COVID-19 consortium
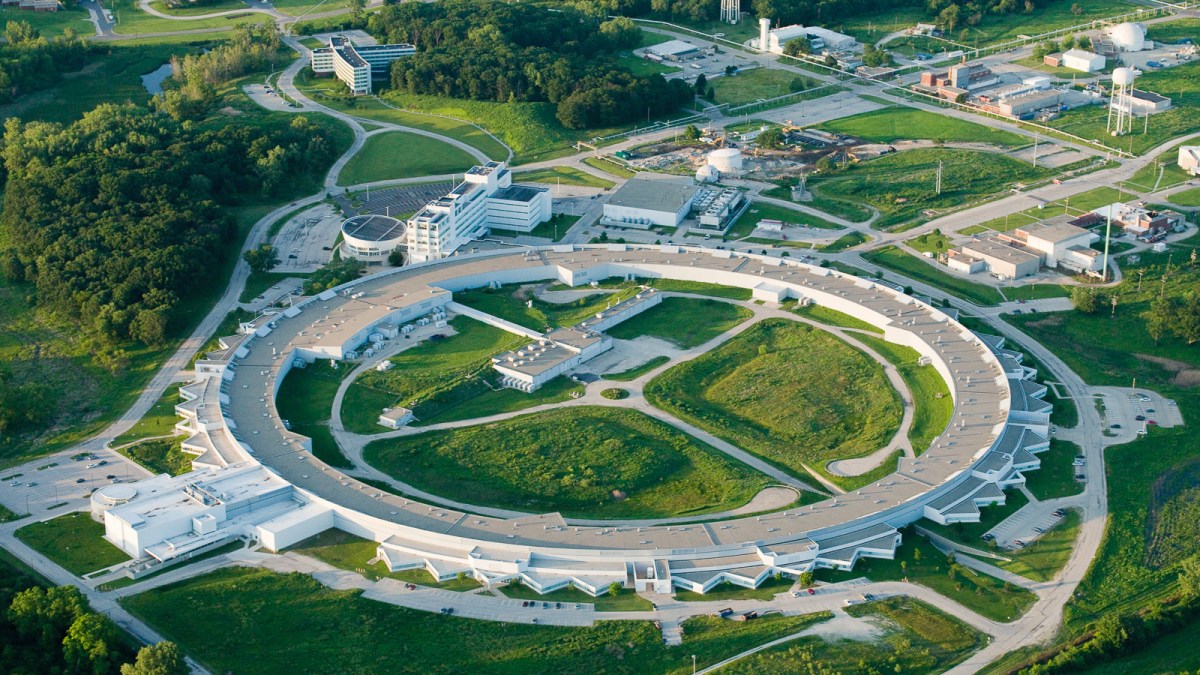

The COVID-19 High Performance Computing Consortium has added another supercomputer to its effort to speed the work of vaccine researchers, with the NVIDIA DGX A100 now operational at Argonne National Laboratory.
The Department of Energy‘s Office of Science is developing a new generation of exascale computers at the Argonne, Oak Ridge and Lawrence national labs, and Argonne is the first DGX A100 buyer, NVIDIA announced Thursday. CEO Jensen Huang says the machine is designed for artificial intelligence applications, “for the end-to-end machine learning workflow — from data analytics to training to inference.”
The compute power “will help researchers explore treatments and vaccines and study the spread of the virus, enabling scientists to do years’ worth of AI-accelerated work in months or days,” said Rick Stevens, an associate lab director at Argonne, in the announcement.
Convened by the White House Office of Science and Technology Policy, the consortium of government, industry and academia already boasts the No. 1 and No. 2 fastest supercomputers in the world: Summit and Sierra.
The DGX A100, meanwhile, has the distinction of being the “fastest AI supercomputer” being used by the consortium, said Kimberly Powell, vice president of health care at NVIDIA, during a Wednesday news briefing. A single node costs $199,000 and delivers 5 petaflops of compute power, and Argonne’s cluster contains 24 nodes — several of which are operational — for 120 petaflops.
For comparison Summit accounts for 200 petaflops, previously 50% of the consortium’s compute power, meaning it remains the fastest supercomputer overall.
Argonne received its system on May 6, began installation May 7 and had it operational May 9, according to a DOE spokesperson.
The DGX A100 can screen 1 billion small molecule drugs in a day, which is “unprecedented” as researchers look for the molecule capable of blocking the coronavirus from binding with cells, Powell said. The faster that drug is identified, the sooner it can be moved into experimentation and clinical trials.
Supercomputers are also being used to assemble genomic data fragments to understand how COVID-19 affects the microbiomes of the lungs and digestive system, and the DGX A100 just helped NVIDIA set a genomics analysis speed record.
“We have just achieved the fastest gold standard in sequencing analysis, and we can sequence a whole genome in less than 20 minutes,” Powell said.
Sequencing the viral genome globally will show researchers how the coronavirus is migrating and mutating, a process that takes NVIDIA seven hours with the help of Oxford Nanopore Technologies’ handheld sequencer. Deployed worldwide, the devices send sequences to a GISAID database of about 25,000 viral genomes.
NVIDIA made its Clara Parabricks software package free to more than 650 COVID-19 researchers now sequencing entire populations.
The tech company also built its own in-house DGX A100 supercomputer, consisting of 50 nodes, to work with the COVID-19 ecosystem in a matter of three weeks.
“That is the beauty of the DGX architecture,” Powell said. “This is a data center-level supercomputer in a node.”